-Search query
-Search result
Showing 1 - 50 of 86 items for (author: dent & k)
EMDB-27770: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 1
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E
EMDB-27771: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 2
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E
EMDB-27772: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 3
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E

EMDB-27773: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 4
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E
EMDB-27774: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 5
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E

PDB-8dxn: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 1
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E

PDB-8dxo: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 2
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E

PDB-8dxp: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 3
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E

PDB-8dxq: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 4
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E

PDB-8dxr: 
Structure of LRRC8C-LRRC8A(IL125) Chimera, Class 5
Method: single particle / : Takahashi H, Yamada T, Denton JS, Strange K, Karakas E

EMDB-15113: 
Human mature large subunit of the ribosome with eIF6 and homoharringtonine bound
Method: single particle / : Faille A, Warren AJ, Dent KC

PDB-8a3d: 
Human mature large subunit of the ribosome with eIF6 and homoharringtonine bound
Method: single particle / : Faille A, Warren AJ, Dent KC

EMDB-14317: 
Cryo-EM reconstruction of the human 40S ribosomal subunit - Full map
Method: single particle / : Pellegrino S, Dent KC, Spikes T, Warren AJ

EMDB-14318: 
Cryo-EM reconstruction of the human 40S ribosomal subunit - Body domain
Method: single particle / : Pellegrino S, Dent KC, Spikes T, Warren AJ

EMDB-14319: 
Cryo-EM reconstruction of the human 40S ribosomal subunit - Head domain
Method: single particle / : Pellegrino S, Dent KC, Spikes T, Warren AJ

PDB-7r4x: 
Cryo-EM reconstruction of the human 40S ribosomal subunit - Full map
Method: single particle / : Pellegrino S, Dent KC, Spikes T, Warren AJ

EMDB-26336: 
APE1 bound to a nucleosome core particle with AP-site at SHL-6
Method: single particle / : Weaver TM, Freudenthal BD
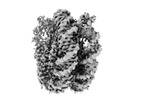
EMDB-26337: 
Nucleosome core particle with AP-site at SHL-6
Method: single particle / : Freudenthal BD, Weaver TM
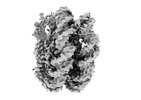
EMDB-26338: 
nucleosome core particle with AP-site at SHL-6.5
Method: single particle / : Freudenthal BD, Weaver TM

EMDB-26339: 
Nucleosome core particle with AP-site at SHL0
Method: single particle / : Freudenthal BD, Weaver TM

PDB-7u50: 
APE1 bound to a nucleosome core particle with AP-site at SHL-6
Method: single particle / : Weaver TM, Freudenthal BD

PDB-7u51: 
Nucleosome core particle with AP-site at SHL-6
Method: single particle / : Freudenthal BD, Weaver TM

PDB-7u52: 
nucleosome core particle with AP-site at SHL-6.5
Method: single particle / : Freudenthal BD, Weaver TM

PDB-7u53: 
Nucleosome core particle with AP-site at SHL0
Method: single particle / : Freudenthal BD, Weaver TM

EMDB-13967: 
Cryo-EM structure of the human mtLSU assembly intermediate upon MRM2 depletion - class 4
Method: single particle / : Rebelo-Guiomar P, Pellegrino S, Dent KC, Warren AJ, Minczuk M

PDB-7qh7: 
Cryo-EM structure of the human mtLSU assembly intermediate upon MRM2 depletion - class 4
Method: single particle / : Rebelo-Guiomar P, Pellegrino S, Dent KC, Warren AJ, Minczuk M

EMDB-13962: 
Cryo-EM structure of the human mtLSU assembly intermediate upon MRM2 depletion - class 2
Method: single particle / : Rebelo-Guiomar P, Pellegrino S, Dent KC, Warren AJ, Minczuk M

EMDB-13963: 
Cryo-EM structure of the human mtLSU assembly intermediate upon MRM2 depletion - class 3
Method: single particle / : Rebelo-Guiomar P, Pellegrino S, Dent KC, Warren AJ, Minczuk M

EMDB-13965: 
Cryo-EM structure of the human mtLSU assembly intermediate upon MRM2 depletion - class 1
Method: single particle / : Rebelo-Guiomar P, Pellegrino S, Dent KC, Warren AJ, Minczuk M

EMDB-13966: 
Cryo-EM structure of the human mtLSU assembly intermediate upon MRM2 depletion - class 5
Method: single particle / : Rebelo-Guiomar P, Pellegrino S, Dent KC, Warren AJ, Minczuk M

PDB-7qh6: 
Cryo-EM structure of the human mtLSU assembly intermediate upon MRM2 depletion - class 1
Method: single particle / : Rebelo-Guiomar P, Pellegrino S, Dent KC, Warren AJ, Minczuk M

EMDB-23066: 
Protective antigen pore translocating lethal factor N-terminal domain
Method: single particle / : Machen AJ, Freudenthal BD

PDB-7kxr: 
Protective antigen pore translocating lethal factor N-terminal domain
Method: single particle / : Machen AJ, Freudenthal BD

EMDB-11692: 
Nsp7-Nsp8-Nsp12 SARS-CoV2 RNA-dependent RNA polymerase in complex with template:primer dsRNA and favipiravir-RTP
Method: single particle / : Naydenova K, Muir KW, Wu LF, Zhang Z, Coscia F, Peet M, Castro-Hartman P, Qian P, Sader K, Dent K, Kimanius D, Sutherland JD, Lowe J, Barford D, Russo CJ

PDB-7aap: 
Nsp7-Nsp8-Nsp12 SARS-CoV2 RNA-dependent RNA polymerase in complex with template:primer dsRNA and favipiravir-RTP
Method: single particle / : Naydenova K, Muir KW, Wu LF, Zhang Z, Coscia F, Peet M, Castro-Hartman P, Qian P, Sader K, Dent K, Kimanius D, Sutherland JD, Lowe J, Barford D, Russo CJ

EMDB-10239: 
Structure of the native full-length HIV-1 capsid protein A92E in helical assembly (-12,11)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

EMDB-10240: 
Structure of the native full-length HIV-1 capsid protein A92E in helical assembly (-13,11)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

EMDB-10246: 
Structure of the native full-length HIV-1 capsid protein in helical assembly (-13,12)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

PDB-6slq: 
Structure of the native full-length HIV-1 capsid protein A92E in helical assembly (-12,11)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

PDB-6slu: 
Structure of the native full-length HIV-1 capsid protein A92E in helical assembly (-13,11)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

PDB-6smu: 
Structure of the native full-length HIV-1 capsid protein in helical assembly (-13,12)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

EMDB-10226: 
Structure of the native full-length HIV-1 capsid protein in helical assembly (-13,8)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

EMDB-10228: 
Structure of the native full-length HIV-1 capsid protein A92E in helical assembly (-13,12)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

EMDB-10229: 
Structure of the native full-length HIV-1 capsid protein in helical assembly (-13,8)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

PDB-6skk: 
Structure of the native full-length HIV-1 capsid protein in helical assembly (-13,8)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

PDB-6skm: 
Structure of the native full-length HIV-1 capsid protein A92E in helical assembly (-13,12)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

PDB-6skn: 
Structure of the native full-length HIV-1 capsid protein in helical assembly (-13,8)
Method: helical / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

EMDB-10738: 
Structure of the native full-length HIV-1 capsid protein in complex with Cyclophilin A from helical assembly (-8,13)
Method: single particle / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

EMDB-10739: 
Structure of the native full-length HIV-1 capsid protein in complex with Cyclophilin A from helical assembly (-13,8)
Method: single particle / : Ni T, Gerard S, Zhao G, Ning J, Zhang P

EMDB-10740: 
Structure of the native full-length HIV-1 capsid protein in complex with Cyclophilin A from helical assembly (-13,7)
Method: single particle / : Ni T, Gerard S, Zhao G, Ning J, Zhang P
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
